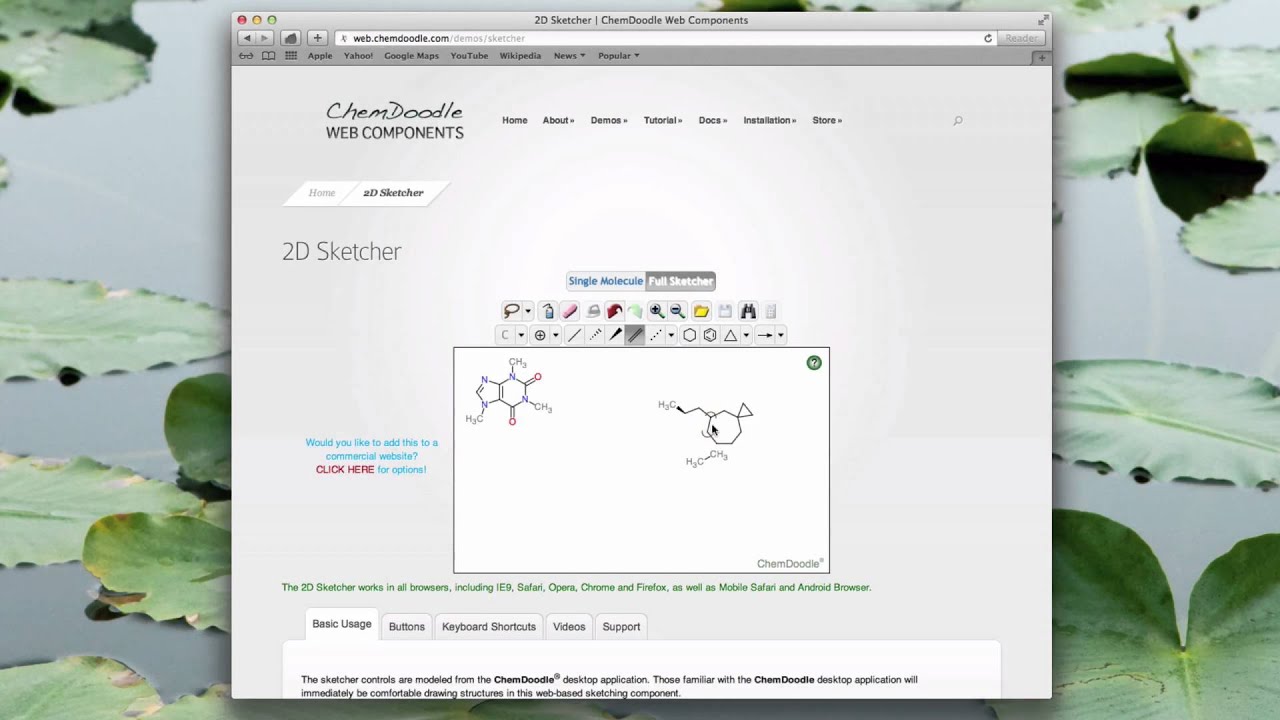 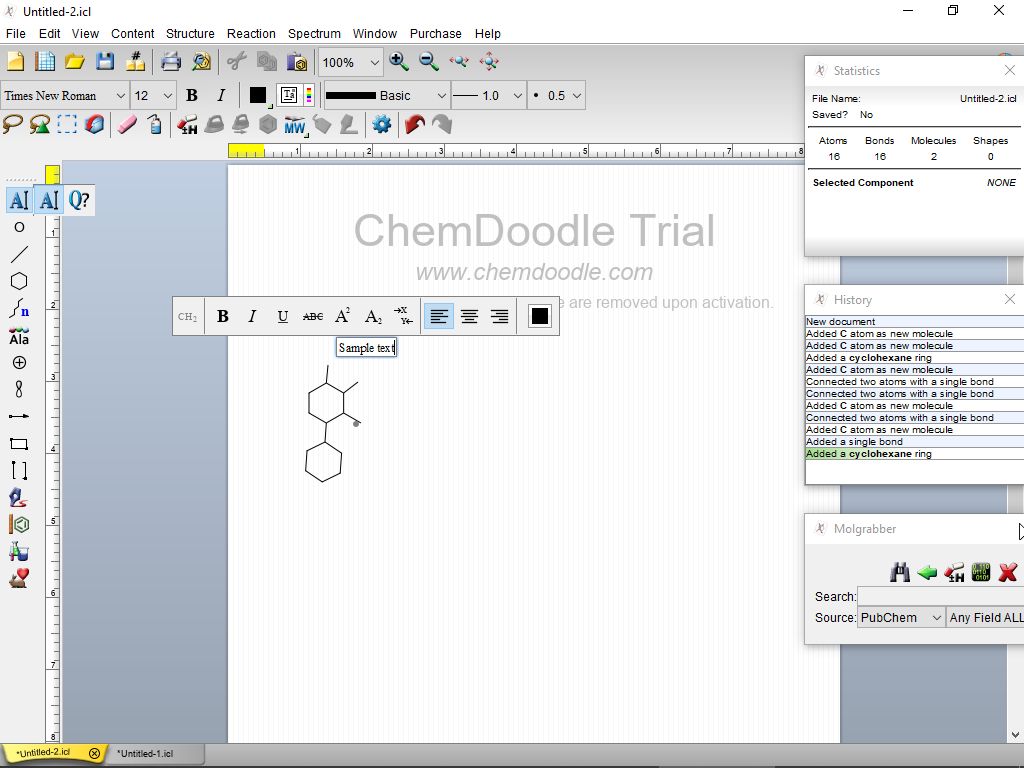
Several times, Microsoft has removed the round-trip editing functionality in Office products. However, there are several issues when relying on round-trip editing, as the user may not have continued access to the originating chemistry program or Microsoft Office, or may be working with an individual on a different operating system. ChemDoodle 2D has provided round-trip editing functionality on Windows, macOS and Linux since its inception and starting with version 10.4, users can paste chemical data from embedded ChemDraw images on macOS in ChemDoodle 2D. Chemists use this all the time when creating chemistry documents in Microsoft Word. This update brings fully chemically aware and self-calculating stoichiometry tables, automatic electron pushing arrows, and with the ability to recover chemical data embedded in Microsoft Office files.Ĭhemical data recovery from Microsoft Office files: (experimental feature) Round-trip editing is the process of embedding data from one application into another, that can then be recovered later. The latest update to the very popular chemical drawing package ChemDoodle has been released. Disable the native file choosers if you are having issues with them, for instance if you have a problem with iCloud on macOS and cannot get the file choosers to load. There is now an option to disable the native file choosers used by ChemDoodle in the Preferences window, under the Files tab. 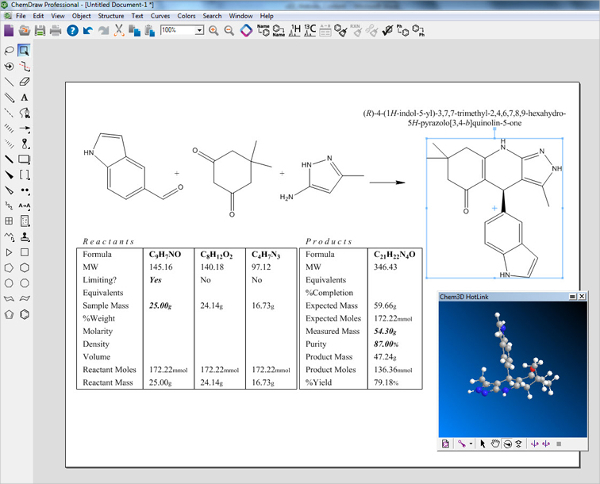 
Updated import of MMTF/PDB data from the RCSB.PubChem access now uses the PUG REST protocol, which results in more stable communication with the PubChem database and higher quality results when using the MolGrabber widget.The SciFinder-n interface has been updated and improved for better usability and with a UI matching the current SciFinder-n product.You may perform structure, substructure and similarity searching into the Google Patents and non-patent literature databases at Google using structures drawn in ChemDoodle. Google Patents searching is integrated.RCSB PDB database connections have been switched to HTTPS. A new PubChem protocol, PUG, replaces the old Entrez protocol for better results from the MolGrabber widget. The SciFinder-n interface has been improved and updated. Google Patents is integrated to allow you to search the Google Patents and non-patent literature databases using structures drawn in ChemDoodle. While minimizations of several discrete structures using through-space forces may look correct, please understand the theory behind the force field you are using and that any force field not specifically developed or parameterized for intermolecular forces will not lead to experimentally accurate coordinates.ChemDoodle 2D v11.5 is a feature update. Please note, while you are able to optimize the entire scene, most force fields are not parameterized for multiple discrete molecular structures, and you will be relying on the through-space forces defined in the force field (mainly van der Waals and electrostatic), if defined at all. To do this, you can change the optimization scope to optimize the entire scene. However, you may wish to optimize several discrete molecular structures at the same time and in relation to each other. ChemDoodle 3D therefore will optimize molecular structures separately and individually as you are editing them. Most small molecule force fields are optimized for describing individual discrete molecular structures.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
